docker build -t transformater .
docker run -it transformater pytest /tests
docker run -it transformater flake8 /transform.py /transformater
docker run -it transformater python transform.py
or pass custom s3 path
docker run -it transformater python transform.py --s3_bucket=backmarket-data-jobs --s3_file_path=data/product_catalog.csv
for large scale testing duplicated and repeated oroginal product calatlog csv to a larger versions:
260mb file
docker run -it transformater python transform.py --s3_bucket=janis-test-bucket --s3_region=eu-central-1 --s3_file_path=product_catalog_260mb.csv
8gb file
docker run -it transformater python transform.py --s3_bucket=janis-test-bucket --s3_region=eu-central-1 --s3_file_path=product_catalog_8gb.csv
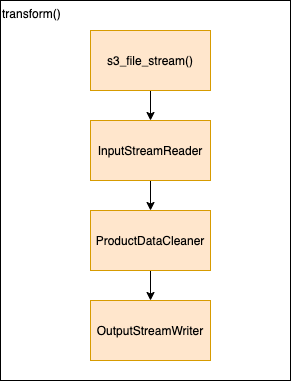
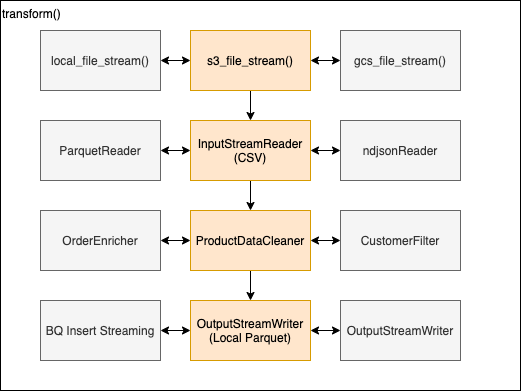
- transform() - orchestration function, iterates over batches and processes each 1 by 1
- s3_file_stream() - open and return s3 file stream without downloading the file
- InputStreamReader - validate schema create batch reader object for CSV and implements batch iterator with retry handling for whhen connection is dropped.
- ProductDataCleaner - performs product filtering for given batch
- OutputStreamWriter - opens output stream and for each batch (valid/invalid) writes and output parquet file
For each of the components code can be extended to support other source, parse and output formats. As well as seperating business logic from generic processing logic easier maintenence.
I chose to write tranformatter project around pyarrow library for 3 reasones:
- support of parquest transofrmations
- support for data streaming
- interoperability for different read/write stream format support like csv, json and filesystem interfaces - local, s3, HDFS
Code is implemented in scalable way and memory profiled. I've tested it with 7GB file it took 10 min to process on my macbook pro with 100 Mbps connection and at peak memory consumption was 200mb. Currently script is built for single process synchronouse processing. Next steps would be to implement it in distributed way.
- Distributed spark infrastructure would be natural next step for large file parallel processing. It would offer little overhead on code implementation side which would be prerequisit in understanding pyspark. And the rest of the complexity would be abstracted by spark infrastructure itself.
- Another alternative would be to use AWS Lambdas or GCP Cloud Functions. In this case I first split the large files in s3 into smaller ones so that each function can process small chunk. Also would need a centralized place for output. such as s3, gcs as storing locally would not be an option. Would be more involved than using spark.
- Leverage k8 infra, same prerequisite - large csv files would need to be split into smaller ones use the docerized version of the script and submit k8 jobs, 1 per file. Same as with Cloud function/Lambdas would require file spliting and centralized output.